Abstract
Object
To enable high-quality physics-guided deep learning (PG-DL) reconstruction of large-scale 3D non-Cartesian coronary MRI by overcoming challenges of hardware limitations and limited training data availability.
Materials and methods
While PG-DL has emerged as a powerful image reconstruction method, its application to large-scale 3D non-Cartesian MRI is hindered by hardware limitations and limited availability of training data. We combine several recent advances in deep learning and MRI reconstruction to tackle the former challenge, and we further propose a 2.5D reconstruction using 2D convolutional neural networks, which treat 3D volumes as batches of 2D images to train the network with a limited amount of training data. Both 3D and 2.5D variants of the PG-DL networks were compared to conventional methods for high-resolution 3D kooshball coronary MRI.
Results
Proposed PG-DL reconstructions of 3D non-Cartesian coronary MRI with 3D and 2.5D processing outperformed all conventional methods both quantitatively and qualitatively in terms of image assessment by an experienced cardiologist. The 2.5D variant further improved vessel sharpness compared to 3D processing, and scored higher in terms of qualitative image quality.
Discussion
PG-DL reconstruction of large-scale 3D non-Cartesian MRI without compromising image size or network complexity is achieved, and the proposed 2.5D processing enables high-quality reconstruction with limited training data.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Non-Cartesian acquisitions are widely utilized in cardiac MRI with their efficient coverage of k-space, less visually apparent aliasing artifacts and higher motion robustness [1,2,3,4]. Among these, 3D non-Cartesian trajectories [5, 6] have been applied in multiple applications, including coronary MRI, for their efficient volumetric coverage [7, 8], and potential to resolve respiratory motion [9,10,11,12] by sampling the center of k-space at every readout. In particular, 3D radial or kooshball trajectories are seldom acquired at Nyquist rate due to the lengthy scan times (about 1.5 times the Nyquist Cartesian acquisition time to achieve the same coverage and resolution), and several accelerated imaging strategies have been studied [9, 13,14,15,16] to shorten the scans, though faster acceleration methods with higher image quality are desirable.
Physics-guided deep learning (PG-DL) has emerged as a powerful reconstruction method for accelerated MRI with improved image quality compared to conventional methods [17,18,19,20,21,22]. A common strategy for PG-DL relies on algorithm unrolling, where an algorithm for solving a regularized least squares problem is unrolled for a fixed number of iterations [17, 20, 21]. The unrolled network alternates between a neural network-based regularizer and a linear data fidelity (DF) unit, which is often solved via gradient descent or conjugate gradient method (CG), and which itself is unrolled. A sufficient number of unrolled iterations, as well as CG steps are required for a satisfactory image quality. Consequently, PG-DL has high GPU memory demands, making its implementation a challenging task for high-resolution MRI, especially with 3D non-Cartesian trajectories, such as kooshball acquisitions [5, 23], where data fidelity cannot be separated across different dimensions.
The difficulties of memory consumption for 3D non-Cartesian MRI are evident in existing studies [24,25,26,27]. In [24] a PG-DL reconstruction of 2D non-Cartesian data was implemented using simulated single-coil acquisitions; in [25] PG-DL of a 3D multi-coil spiral acquisition was performed for low-resolution navigator data. On the other hand, alternative strategies that alter the model formulation have also been proposed: In [26] a 3D Cartesian volume was broken into smaller 3D slabs along frequency encode dimension to fit the GPU memory; while [27] proposed block-wise DF using limited CG iterations to reduce GPU memory consumption. An additional issue for implementing PG-DL reconstruction for 3D non-Cartesian trajectories is the limited availability of training data. In this case, each imaging volume corresponds to a single subject, and data augmentation strategies such as the one in [26] are not applicable.
In this study, we combined several recent advances in DL and MRI reconstruction to enable PG-DL reconstruction of multi-coil 3D kooshball coronary MRI data. From an implementation perspective, we combined recently developed memory-efficient learning (MEL) techniques [28], the Toeplitz formulation for the operator used in DF units [29, 30], mixed-precision processing [31] and distributed learning [32] to ensure deep networks can be implemented without restrictions on the architectures. Furthermore, to tackle the challenge of limited training data, we proposed a 2.5D strategy that efficiently uses the 3D training data with a regularization process that comprises three 2D convolutional neural networks (CNNs) in orthogonal views. Our results suggest that both the 3D and 2.5D PG-DL implementations outperform conventional linear reconstructions [33, 34], with the 2.5D strategy leading to higher vessel sharpness and improved qualitative image scores.
Materials and methods
Physics-Guided deep learning formulation
PG-DL solves the regularized multi-coil reconstruction model:
where \(\mathbf{y}\) is acquired k-space, \(\mathbf{E}\) is multi-coil encoding operator, \(\mathbf{x}\) is the underlying image, and \(\mathcal{R}\left(\cdot \right)\) is a regularizer. In PG-DL, the objective function in (1) can be solved with various algorithms [35], including variable splitting with quadratic penalty [17, 19, 21]:
where \(\mu\) is a learnable scalar. The proximal operator for the regularizer in Eq. (2) is solved by a trained CNN-based regularizer, followed by the data fidelity (DF) step in Eq. (3) whose solution can be estimated via conjugate gradient (CG) method [19, 21]. The reconstruction is given by a pre-determined number of unrolled iterations between Eqs. (2) and (3) (Fig. 1). Since the DF term performs multiple CG iterations itself, the resulting computational graph may lead to large GPU memory demands for backpropagation.
PG-DL scheme for 3D high-resolution multi-coil non-Cartesian MRI. a PG-DL starts from a linear reconstruction with one CG iteration. MEL is applied for the unrolled network, to support unlimited unrolled iterations using GPU memory of a single unrolled step. The CNN regularizer is implemented using half-precision fitting into a single GPU. b In MEL, intermediate data in the forward propagation is stored in host memory and the GPU resource is released and reused for the next unrolled iteration. During backpropagation, the intermediate results on host memory is fed to the GPU to the gradients are computed through each unrolled step. c CG iterations in DF is performed on an individual GPU, which distributes coil images to additional GPUs during multi-coil encoding and decoding operations. DF is implemented using single precision to support k-space processing
Memory efficient learning for algorithm unrolling
The alternating feedforward structure of the unrolled network suggests that each unrolled iteration can be performed sequentially. To this end, an MEL scheme for PG-DL was proposed in [28], which reduces GPU memory consumption of a PG-DL network with \(N\) unrolled iterations from \(\mathcal{O}(N)\) to \(\mathcal{O}(1)\), with trade-offs between increased computational complexity for lower memory consumption. The scheme in [28] was a special case of the broader technique of memory checkpointing [36,37,38], which builds the computation graph as a single unrolled step on the device (GPU), and save the intermediate outputs on the host (CPU) memory (Fig. 2a). During forward propagation, the output images of each unrolled step is saved on host memory, to be used for computing gradient of each unrolled step during backpropagation, where the gradient is also saved step by step on the host memory. The resulting PG-DL scheme uses GPU memory corresponding to a single unrolled step while supporting an arbitrary number of unrolling steps (Fig. 1b).
Toeplitz method for data fidelity
The DF step (Eq. 3) computes \({\mathbf{E}}^{H}\mathbf{E}\), which consists of NUFFT and its adjoint operation, in every CG iteration. This requires large GPU memory consumption during backpropagation, especially with respect to the convolutions in NUFFT, and often prevents the training of PG-DL for 3D multi-coil non-Cartesian MRI. The Toeplitz nature of \({\mathbf{E}}^{H}\mathbf{E}\) has been noted [29, 30], and has been used to convert this calculation into a series of fast Fourier transform and point-wise multiplications with a trajectory-dependent operator on a zero-padded image as:
where \({\mathbf{S}}_{n}\) denotes the \(n\)th coil sensitivities, \({n}_{c}\) is the number of coils, \(\mathbf{Z}\) is zero-padding to double FOV along each dimension and \({\mathbf{Z}}^{H}\) is cropping to the original FOV, \(\mathbf{F}\) is Cartesian FFT, and \(\mathbf{M}\) is a trajectory-dependent operator that can be pre-computed as:
where \({\mathcal{F}}^{H}\) denotes adjoint NUFFT that grids non-Cartesian k-space of doubled FOV along each dimension, \(\mathbf{W}\) is density compensation weights, and \({\varvec{\updelta}}\) is the delta impulse function. Under this setup, the non-Cartesian PG-DL is implemented without the use of NUFFT or its adjoint, but by Cartesian FFTs and diagonal matrix multiplication, leading to substantial savings in computational time.
Mixed precision
Half-precision processing uses 16-bit floating point data (float16) that naturally occupies less memory than conventional single precision (float32). Although float16 has a limited representation range, its application in computer vision is well-established with virtually no impact on network accuracy from the use of half-precision [31]. However, unlike images used in computer vision, k-space data exhibit large variations of intensities hindering the use of half-precision. Thus, in this study, we apply half-precision for the CNN regularizer only, while the DF employs conventional single precision (float32) for a sufficient representable range. As a result, the depth of the CNN regularizer can be high while only requiring access to a single GPU, and the DF unit does not incur any risks of over/underflow.
Distributed learning
The zero-padding in the Toeplitz method increases the image matrix size by 23 = 8 times for 3D data. Additionally, as aforementioned, k-space operations are not well-suited for half-precision processing. Thus, considerable GPU memory is required to support multi-coil DF units. Noting the separability of Eq. (4) across coils, we distribute the operation DF to multiple devices. Based on the GPU configuration, we split all \({n}_{c}\) coils into three subsets to be evaluated on three GPUs respectively, and combine the results in a primal device (Fig. 1c).
2.5D PG-DL for 3D reconstruction
To train the PG-DL network with limited training data, we propose a 2.5D PG-DL formulation using 3D DF and three 2D CNN regularizers \({\mathcal{R}}_{c}(\cdot )\), \({\mathcal{R}}_{s}(\cdot )\), \({\mathcal{R}}_{a}(\cdot )\) over coronal, sagittal, and axial planes, respectively:
Using the same variable splitting with quadratic penalty approach, we have:
which is solved via:
where the proximal operators in (8)–(10) are solved by three distinct 2D CNNs, and the DF term in (11) is solved via CG as before. The 2D CNNs take 3D volume as input while treating the direction orthogonal to the plane of interest as the batch dimension, thus each 3D volume provides hundreds of 2D images for training (Fig. 2).
Coronary MRI datasets
3D kooshball coronary MRI was acquired on a 1.5 T scanner (Magnetom Aera, Siemens Healthcare, Erlangen, Germany) on 9 subjects, using an ECG-triggered T2-prepared, fat-saturated, navigator-gated bSSFP sequence, with resolution = (1.15 mm)3, matrix size = 1923 (grid size = 3843), FOV = (220 mm)3 and twofold readout oversampling. A total of 12,320 radial projections (sub-Nyquist rate of 5) were acquired in 385 heartbeats using the spiral phyllotaxis pattern [23] with one interleaf of 32 readouts per heartbeat, resulting in scan efficiency of 41.16 ± 11.75 (%). The data was further retrospectively sixfold undersampled prior to any processing.
Implementation details
As detailed earlier, the CNN-based regularizer is implemented using half-precision to fit a ResNet [21] with sufficient depth on a single GPU. The proposed 3D PG-DL uses a ResNet regularizer comprising 10 residual blocks, where each residual block has 2 convolutional layers using 3 × 3 × 3 convolutions without bias, followed by ReLU activation. Such ResNet results in 1,444,611 learnable parameters. 2.5D PG-DL employs three 2D ResNets of the same architecture as its 3D counterpart, only replacing the 3D convolutions with 3 × 3 2D convolutions. In such case, three 2D ResNets lead to the same number of learnable parameters as a 3D ResNet using 3 × 3 × 3 convolutions. Both 3D and 2.5D PG-DL has 10 unrolled iterations and 10 CG iterations in the DF. DF is implemented using single precision on an individual GPU, which further distributes coil images into additional 2 GPUs to perform the \({\mathbf{E}}^{H}\mathbf{E}\) operations. All experiments were implemented using Pytorch 1.14, and executed on a server with 4 NVIDIA A100 GPUs (40 GB). PG-DL networks were trained using an ADAM optimizer with a learning rate of 0.0005 for 100 epochs. A mixed \({{\ell}}_{1}\)–\({{\ell}}_{2}\) loss was employed [21] to perform supervised training using 5 of 18 subjects as training data, and the remaining 13 distinct subjects were used for testing. For each subject, a 3D volume is used for the 3D PG-DL, while 2.5D PG-DL takes 192 2D slices chosen equally from the 3 orthogonal views. The peak memory consumption during training were 107.1 GB and 74.0 GB for 3D and 2.5D set-up respectively, and 79.4 GB and 46.1 GB for 3D and 2.5D, respectively during testing. For comparison, the datasets were also reconstructed using conventional gridding and Tikhonov-regularized CG-SENSE reconstruction.
Image analysis
Quantitative assessments using SSIM and PSNR were performed with respect to the fully-sampled reference image. Vessel sharpness scores and vessel length were obtained using SoapBubble software [39]. Since CG-SENSE and PG-DL outputs complex coil-combined images as their final results, we adapted complex coil combination [40, 41] to generate the images for a fair comparison between reference and other reconstructions, instead of root-sum-squares combination [23, 42]. Statistical differences in SSIM, PSNR and vessel sharpness scores were assessed using a paired signed rank test with a P value < 0.05 being considered significant.
Furthermore, a qualitative image quality assessment was performed by an experienced cardiologist (15 years of experience). The reader was blinded to the reconstruction methods, and their orders were randomized. Four test subjects were evaluated on a 4-point ordinal scale, for overall image quality (1: excellent; 2: good; 3: fair; 4: poor), perceived SNR (1: high SNR; 2: minor noise with moderate SNR; 3: major noise but not limiting clinical diagnosis; 4: poor SNR and non-diagnostic), aliasing artifacts (1: no visual apparent aliasing; 2: mild; 3: moderate; 4: severe) and blurring (1: none; 2: mild; 3: moderate; 4: severe).
Results
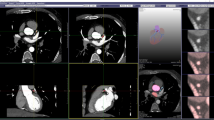
Figure 3 depicts representative reconstruction results of retrospectively sixfold undersampled data, using gridding, Tikhonov-regularized CG-SENSE, 3D PG-DL and 2.5D PG-DL. Due to the relatively high undersampling rate, gridding reconstruction exhibited blurring and undersampling artifacts. Tikhonov-regularized CG-SENSE showed less blurring than gridding but suffers from noise amplification due to the high acceleration rate. Both 3D and 2.5D PG-DL substantially reduced the noise compared to Tikhonov regularized CG-SENSE, while 2.5D PG-DL offered visually improved image sharpness compared to its 3D counterpart.
Representative reconstruction results with retrospectively undersampled data at rate 6, using gridding, Tikhonov regularized CGSENSE, 3D PG-DL and 2.5D PG-DL. Gridding exhibits blurring and undersampling artifacts. Tikhonov regularized CGSENSE displays noise amplification and blurring. Both 3D and 2.5D PG-DL offer reduced noise and artifacts, while 2.5D PG-DL outperforms 3D PG-DL in terms of sharpness
Figure 4 depicts the quantitative metrics of the tested reconstruction methods. Boxplots of SSIM (Fig. 4a) and NMSE (Fig. 4b) suggest the gridding led to the lowest quantitative metrics due to the relatively high undersampling rate. Tikhonov-regularized CG-SENSE improved SSIM and NMSE compared to gridding, while both 3D and 2.5D PG-DL achieved significantly better scores (P < 0.05). No statistical difference were observed between 3D and 2.5D PG-DL. Similar conclusions apply to vessel sharpness measurements (Fig. 4c) with gridding having the lowest sharpness in all subjects, CGSENSE outperforming gridding with limited improvement, and 2.5D PG-DL and 3D PG-DL substantially improving on these methods. In particular, 2.5D PG-DL had higher vessel sharpness in all subjects compared to 3D PG-DL, performing closely to fully-sampled reference. Similar to PSNR and SSIM, 2.5D and 3D PG-DL significantly improved upon classical methods, though the differences among each other were not significant. Measured vessel length (Fig. 4d) results suggest gridding suffered from significant sub-sampling artifacts and failed to provide reasonable vessel length; both 3D and 2.5D PG-DL gave more accurate vessel length measurements than Tikhonov-regularized CG-SENSE, while 2.5D PG-DL shows an advantage over its 3D counterpart.
Quantitative metrics of tested methods. a SSIM computed on 2D slices in the 3 orthogonal views of 13 testing subjects. Medians are 0.7402, 0.7770, 0.8412, 0.8441 for gridding, Tikhonov-regularized CGSENSE, 3D PG-DL and 2.5D PG-DL, respectively. b PSNR of the tested methods, computed from 2D slices in the three orthogonal views of all testing data with medians 22.5034, 30.4560, 32.1610 and 32.2839 for gridding, Tikhonov-regularized CGSENSE, 3D PG-DL and 2.5D PG-DL, respectively. c Vessel sharpness scores (%) of the reference fully sampled image and tested methods with medians 76.1980, 38.7044, 63.3126, 74.3423 and 76.7669 for reference, gridding, CGSENSE, 3D and 2.5D PG-DL, respectively. 2.5D PG-DL and 3D PG-DL had similar SSIM and NMSE, while 2.5D PG-DL gave higher sharpness scores than 3D PG-DL in all tested subjects. d Measured vessel length of the fully sampled image and tested methods for all tested subjects, with averaged vessel length of 4.8187 ± 1.1595 cm, 4.562 ± 0. 6267 cm, 4.6477 ± 0. 8476 cm, 4.7911 ± 1. 1259 and 4.8039 ± 1.1465 cm, for gridding, Tikhonov-regularized CGSENSE, 3D PG-DL and 2.5D PG-DL respectively, with respect to the fully sampled image. Both PG-DL gave more accurate vessel length measurement, where 2.5D PG-DL showed lower error than its 3D counterpart
Figure 5 depicts the reader study results for all methods. 2.5D PG-DL gives excellent overall image quality. Both 3D and 2.5D PG-DL show high SNR, no visual apparent aliasing and mild blurring. CG-SENSE shows moderate to severe blur while gridding suffers from low SNR and severe blurring.
Clinical reader study results of tested methods. A 4-point ordinal scale (1: excellent, 2: good, 3: fair, 4: poor) was employed, for overall image quality, perceived SNR, aliasing artifacts and blurring. 2.5D PG-DL gave an overall image quality similar to the reference image. Both 2.5D and 3D PG-DL showed high SNR, no visual aliasing and mild blurring, outperforming gridding and CG-SENSE
Discussion
In this study, we investigated PG-DL of high-resolution, multi-coil non-Cartesian coronary MRI from a practical perspective. In addition to several memory-efficient techniques to tackle hardware limitations, a 2.5D processing strategy was proposed to improve PG-DL using limited training data. Our results show that PG-DL significantly improved image quality compared to conventional methods, while 2.5D PG-DL further led to sharper and higher-rated image quality compared to 3D PG-DL in this limited training data setup.
The combination of MEL, Toeplitz method and distributed learning is necessary in training to ensure a sufficient number of unrolled iterations as well as CG iterations for reconstruction. Without using MEL, only 1 unrolled iteration would be supported using the 4-GPU system with a total memory of 160 GB. In a preliminary study [43], we showed that half-precision was needed to ensure a ResNet with sufficient depth, and distributed learning was necessary to facilitate the use of all 32 coils in DF without coil compression.
The Toeplitz method was first proposed to accelerate non-Cartesian reconstruction [29, 30] by trading memory (8 times for 3D case) with processing speed. Note it has the same memory consumption as NUFFT with 2 × oversampling. However, the Toeplitz method benefits the PG-DL training by simplifying backpropagation using point-wise multiplications instead of convolutions. As a result, PG-DL using the Toeplitz method took less memory than using NUFFT with 1.25 × oversampling during the training. Even with the use of MEL, we failed to execute the training using NUFFT [44] for DF instead of the Toeplitz method due to higher memory demands.
Existing studies on 2.5D networks focus on its close performance, faster processing speed and lower memory consumption compared to 3D networks [45,46,47,48]. These studies, mostly confined to computer vision applications investigate potentially better performance than 3D networks under memory constrains. In our work, the proposed 2.5D PG-DL is formulated via 3D DF and 2D regularization terms, where view-aggregating is achieved via quadratic relaxation with learnable regularization strength. As reported in [48], 2.5D networks with view-aggregating outperform 3D networks using constrained resources, while 3D networks are believed to eventually outperform 2.5D given unlimited resources, for their ability to better model 3D textures [48]. As reported in [48], 2.5D networks with view-aggregating outperform 3D networks using constrained resources, while 3D networks are believed to eventually outperform 2.5D given unlimited resources, for their ability to better model 3D textures [48]. Due to the lack of sufficient training data (in total 5 volumes), it is unclear if 3D PG-DL can eventually outperform 2.5D PG-DL given sufficient data during the training, in the context of MRI reconstruction. In this study, the 2.5D processing allowed improved PG-DL networks to be trained from a higher number of 2D slices instead of limited 3D volumes. As the limited training data becomes an additional challenge for 3D non-Cartesian PG-DL, the 2.5D formulation provides a feasible strategy to exploit the training data efficiently, without introducing artificial augmentation strategies which may be insufficient with such limited training subjects [49]. Finally, we note that while the improvements between 2.5D PG-DL and 3D PG-DL were not statistically significant, this may be due to the limited size of the test database.
In this study, we have used normalized vessel sharpness, as determined by SoapBubble software [39] to evaluate the sharpness of the reconstructions, as is commonly done in coronary MRI studies [13, 50, 51]. However, we note that there are other edge sharpness measurements that have been proposed for cardiac MRI that may also be useful in other applications [52].
This study focused on enabling PG-DL reconstruction under two major restrictions of GPU memory and limited training data, and building a first-of-its-kind baseline for PG-DL reconstruction of large-sized 3D non-Cartesian MRI data. To this end, a combination of techniques was used to ensure the results were compatible with commonly used techniques in the field. Despite the proposed combination of memory-efficient techniques, several recent advancements are worthy to be considered in future works. For model unrolling, deep equilibrium models [53, 54] are an alternative method that offers a memory-efficient strategy for supporting arbitrary numbers of unrolled iterations that are run until convergence. However, many existing studies in the field use a fixed number of unrolled steps, thus we adapted MEL to make the framework compatible with the commonly used reconstruction techniques. Although the Toeplitz method costs less memory than NUFFTs, it significantly increases the matrix size during DF. It is worthy to investigate memory-efficient approximations to replace gridding and its adjoint operation, such as [27, 55, 56]. However, all these compression-type techniques cause error accumulation through unrolls. Thus, in this study we adopted the Toeplitz method without coil compression as a baseline. To handle insufficient amount of training data, transfer learning approaches have also been suggested [57, 58]. However, the performance of transfer learning is largely dependent on the prior task from which the transfer is made. Thus, it was not considered in this work but may be worth investigating in future studies. In conclusion, PG-DL of 3D high-resolution multi-coil non-Cartesian coronary MRI is achieved in this study, without compromising multi-coil data size or network depth. We further formulated a 2.5D PG-DL for 3D reconstruction to efficiently utilize limited training data, and obtained visually improved image quality compared to the conventional 3D PG-DL.
Data availability
In accordance with the institutional review board, the data acquired in this study contain person-sensitive information, and can only be shared in the context of scientific collaborations.
References
Thedens DR, Irarrazaval P, Sachs TS, Meyer CH, Nishimura DG (1999) Fast magnetic resonance coronary angiography with a three-dimensional stack of spirals trajectory. Magn Reson Med 41(6):1170–1179
Wright KL, Hamilton JI, Griswold MA, Gulani V, Seiberlich N (2014) Non-Cartesian parallel imaging reconstruction. J Magn Reson Imaging 40(5):1022–1040
Chen Y, Lo WC, Hamilton JI, Barkauskas K, Saybasili H, Wright KL et al (2018) Single breath-hold 3D cardiac T1 mapping using through-time spiral GRAPPA. NMR Biomed 31(6):e3923
Irarrazabal P, Nishimura DG (1995) Fast three dimensional magnetic resonance imaging. Magn Reson Med 33(5):656–662
Stehning C, Börnert P, Nehrke K, Eggers H, Dössel O (2004) Fast isotropic volumetric coronary MR angiography using free-breathing 3D radial balanced FFE acquisition. Magn Reson Med 52(1):197–203
Gurney PT, Hargreaves BA, Nishimura DG (2006) Design and analysis of a practical 3D cones trajectory. Magn Reson Med 55(3):575–582
Barger AV, Block WF, Toropov Y, Grist TM, Mistretta CA (2002) Time-resolved contrast-enhanced imaging with isotropic resolution and broad coverage using an undersampled 3D projection trajectory. Magn Reson Med 48(2):297–305
Feng L, Grimm R, Block KT, Chandarana H, Kim S, Xu J et al (2014) Golden-angle radial sparse parallel MRI: combination of compressed sensing, parallel imaging, and golden-angle radial sampling for fast and flexible dynamic volumetric MRI. Magn Reson Med 72(3):707–717
Piccini D, Feng L, Bonanno G, Coppo S, Yerly J, Lim RP et al (2017) Four-dimensional respiratory motion-resolved whole heart coronary MR angiography. Magn Reson Med 77(4):1473–1484
Bonanno G, Piccini D, Marchal B, Zenge M, Stuber M (2014) A new binning approach for 3D motion corrected self-navigated whole-heart coronary MRA using independent component analysis of individual coils. In: Proceedings of the 22nd annual meeting of ISMRM, Milan, p 936
Larson AC, White RD, Laub G, McVeigh ER, Li D, Simonetti OP (2004) Self-gated cardiac cine MRI. Magn Reson Med 51(1):93–102
Stehning C, Börnert P, Nehrke K, Dössel O (2005) Free breathing 3D balanced FFE coronary magnetic resonance angiography with prolonged cardiac acquisition windows and intra-RR motion correction. Magn Reson Med 53(3):719–723
Feng L, Coppo S, Piccini D, Yerly J, Lim RP, Masci PG et al (2018) 5D whole-heart sparse MRI. Magn Reson Med 79(2):826–838
Mistretta CA, Wieben O, Velikina J, Block W, Perry J, Wu Y et al (2006) Highly constrained backprojection for time-resolved MRI. Magn Reson Med 55(1):30–40
Feng L, Delacoste J, Smith D, Weissbrot J, Flagg E, Moore WH et al (2019) Simultaneous evaluation of lung anatomy and ventilation using 4D respiratory-motion-resolved ultrashort echo time sparse MRI. J Magn Reson Imaging 49(2):411–422
Nam S, Akcakaya M, Basha T, Stehning C, Manning WJ, Tarokh V et al (2013) Compressed sensing reconstruction for whole-heart imaging with 3D radial trajectories: a graphics processing unit implementation. Magn Reson Med 69(1):91–102
Hammernik K, Klatzer T, Kobler E, Recht MP, Sodickson DK, Pock T et al (2018) Learning a variational network for reconstruction of accelerated MRI data. Magn Reson Med 79(6):3055–3071
Schlemper J, Caballero J, Hajnal JV, Price A, Rueckert D (2017) A deep cascade of convolutional neural networks for MR image reconstruction. In: Information processing in medical imaging: 25th international conference, IPMI 2017, Boone, NC, USA, June 25–30, 2017, Proceedings 25. Springer, pp 647–658
Aggarwal HK, Mani MP, Jacob M (2018) MoDL: Model-based deep learning architecture for inverse problems. IEEE Trans Med Imaging 38(2):394–405
Knoll F, Hammernik K, Zhang C, Moeller S, Pock T, Sodickson DK et al (2020) Deep-learning methods for parallel magnetic resonance imaging reconstruction: a survey of the current approaches, trends, and issues. IEEE Signal Process Mag 37(1):128–140
Yaman B, Hosseini SAH, Moeller S, Ellermann J, Uğurbil K, Akçakaya M (2020) Self-supervised learning of physics-guided reconstruction neural networks without fully sampled reference data. Magn Reson Med 84(6):3172–3191
Hammernik K, Küstner T, Yaman B, Huang Z, Rueckert D, Knoll F et al (2023) Physics-driven deep learning for computational magnetic resonance imaging: combining physics and machine learning for improved medical imaging. IEEE Signal Process Mag 40(1):98–114
Piccini D, Littmann A, Nielles-Vallespin S, Zenge MO (2011) Spiral phyllotaxis: the natural way to construct a 3D radial trajectory in MRI. Magn Reson Med 66(4):1049–1056
Ramzi Z, Chaithya G, Starck J-L, Ciuciu P (2022) NC-PDNet: a density-compensated unrolled network for 2D and 3D non-Cartesian MRI reconstruction. IEEE Trans Med Imaging 41(7):1625–1638
Malavé MO, Baron CA, Koundinyan SP, Sandino CM, Ong F, Cheng JY et al (2020) Reconstruction of undersampled 3D non-Cartesian image-based navigators for coronary MRA using an unrolled deep learning model. Magn Reson Med 84(2):800–812
Deng Z, Yaman B, Zhang C, Moeller S, Akçakaya M (2021) Efficient training of 3D unrolled neural networks for MRI reconstruction using small databases. In: 2021 55th Asilomar conference on signals, systems, and computers. IEEE, pp 886–889.
Chen Z, Chen Y, Xie Y, Li D, Christodoulou AG (2022) Data-consistent non-Cartesian deep subspace learning for efficient dynamic MR image reconstruction. In: 2022 IEEE 19th international symposium on biomedical imaging (ISBI). IEEE, pp 1–5
Kellman M, Zhang K, Markley E, Tamir J, Bostan E, Lustig M et al (2020) Memory-efficient learning for large-scale computational imaging. IEEE Trans Comput Imaging 6:1403–1414
Baron CA, Dwork N, Pauly JM, Nishimura DG (2018) Rapid compressed sensing reconstruction of 3D non-Cartesian MRI. Magn Reson Med 79(5):2685–2692
Ramani S, Fessler JA (2013) Accelerated nonCartesian SENSE reconstruction using a majorize-minimize algorithm combining variable-splitting. In: 2013 IEEE 10th international symposium on biomedical imaging. IEEE, pp 704–707
Micikevicius P, Narang S, Alben J, Diamos G, Elsen E, Garcia D et al (2017) Mixed precision training. arXiv:1710.03740
Zhou L, Pan S, Wang J, Vasilakos AV (2017) Machine learning on big data: opportunities and challenges. Neurocomputing 237:350–361
Pruessmann KP, Weiger M, Scheidegger MB, Boesiger P (1999) SENSE: sensitivity encoding for fast MRI. Magn Reson Med 42(5):952–962
Pruessmann KP, Weiger M, Börnert P, Boesiger P (2001) Advances in sensitivity encoding with arbitrary k-space trajectories. Magn Reson Med 46(4):638–651
Fessler JA (2020) Optimization methods for magnetic resonance image reconstruction: Key models and optimization algorithms. IEEE Signal Process Mag 37(1):33–40
Kumar R, Purohit M, Svitkina Z, Vee E, Wang J (2019) Efficient rematerialization for deep networks. In: Advances in neural information processing systems, p 32
Griewank A (1999) An implementation of checkpointing for the reverse or adjoint model of differentiation. ACM Trans Math Softw 26(1):1–19
Jain P, Jain A, Nrusimha A, Gholami A, Abbeel P, Gonzalez J et al (2020) Checkmate: breaking the memory wall with optimal tensor rematerialization. Proc Mach Learn Syst 2:497–511
Etienne A, Botnar RM, Van Muiswinkel AM, Boesiger P, Manning WJ, Stuber M (2002) “Soap-Bubble” visualization and quantitative analysis of 3D coronary magnetic resonance angiograms. Magn Reson Med 48(4):658–666
Hutchinson M, Raff U (1988) Fast MRI data acquisition using multiple detectors. Magn Reson Med 6(1):87–91
Ra JB, Rim C (1993) Fast imaging using subencoding data sets from multiple detectors. Magn Reson Med 30(1):142–145
Cruz G, Atkinson D, Henningsson M, Botnar RM, Prieto C (2017) Highly efficient nonrigid motion-corrected 3D whole-heart coronary vessel wall imaging. Magn Reson Med 77(5):1894–1908
Zhang C, Piccini D, Demirel OB, Bonanno G, Yaman B, Stuber M et al (2022) Distributed memory-efficient physics-guided deep learning reconstruction for large-scale 3d non-Cartesian MRI. In: 2022 IEEE 19th international symposium on biomedical imaging (ISBI). IEEE, pp 1–5
Muckley MJ, Stern R, Murrell T, Knoll F (2020) TorchKbNufft: a high-level, hardware-agnostic non-uniform fast Fourier transform. In: ISMRM workshop on data sampling & image reconstruction
Minnema J, Wolff J, Koivisto J, Lucka F, Batenburg KJ, Forouzanfar T et al (2021) Comparison of convolutional neural network training strategies for cone-beam CT image segmentation. Comput Methods Programs Biomed 207:106192
Bermejo-Peláez D, Estepar RSJ, Ledesma-Carbayo MJ (2018) Emphysema classification using a multi-view convolutional network. In: 2018 IEEE 15th international symposium on biomedical imaging (ISBI 2018). IEEE, pp 519–522
Ziabari A, Ye DH, Srivastava S, Sauer KD, Thibault J-B, Bouman CA (2018) 2.5 D deep learning for CT image reconstruction using a multi-GPU implementation. In: 2018 52nd Asilomar conference on signals, systems, and computers. IEEE, pp 2044–2049
Prasoon A, Petersen K, Igel C, Lauze F, Dam E, Nielsen M (2013) Deep feature learning for knee cartilage segmentation using a triplanar convolutional neural network. In: Medical image computing and computer-assisted intervention–MICCAI 2013: 16th international conference, Nagoya, Japan, September 22–26, 2013, Proceedings, Part II 16. Springer, pp 246–253
Shorten C, Khoshgoftaar TM (2019) A survey on image data augmentation for deep learning. J Big Data 6(1):1–48
Henningsson M, Koken P, Stehning C, Razavi R, Prieto C, Botnar RM (2012) Whole-heart coronary MR angiography with 2D self-navigated image reconstruction. Magn Reson Med 67(2):437–445
Küstner T, Munoz C, Psenicny A, Bustin A, Fuin N, Qi H et al (2021) Deep-learning based super-resolution for 3D isotropic coronary MR angiography in less than a minute. Magn Reson Med 86(5):2837–2852
Ahmad R, Ding Y, Simonetti OP (2015) Edge sharpness assessment by parametric modeling: application to magnetic resonance imaging. Concepts Magn Reson Part A 44(3):138–149
Gilton D, Ongie G, Willett R (2021) Deep equilibrium architectures for inverse problems in imaging. IEEE Trans Comput Imaging 7:1123–1133
Güngör A, Askin B, Soydan DA, Top CB, Saritas EU, Çukur T (2023) DEQ-MPI: a deep equilibrium reconstruction with learned consistency for magnetic particle imaging. IEEE Trans Med Imaging
Seiberlich N, Breuer F, Blaimer M, Jakob P, Griswold M (2008) Self-calibrating GRAPPA operator gridding for radial and spiral trajectories. Magn Reson Med 59(4):930–935
Akçakaya M, Nam S, Basha TA, Kawaji K, Tarokh V, Nezafat R (2014) An augmented Lagrangian based compressed sensing reconstruction for non-Cartesian magnetic resonance imaging without gridding and regridding at every iteration. PLoS ONE 9(9):e107107
Yurt M, Özbey M, Dar SU, Tinaz B, Oguz KK, Çukur T (2022) Progressively volumetrized deep generative models for data-efficient contextual learning of MR image recovery. Med Image Anal 78:102429
Dar SUH, Özbey M, Çatlı AB, Çukur T (2020) A transfer-learning approach for accelerated MRI using deep neural networks. Magn Reson Med 84(2):663–685
Funding
NIH, Grant numbers: R01HL153146, R01EB032830, R21EB028369, P41EB027061.
Author information
Authors and Affiliations
Contributions
Mehmet Akçakaya: Study conception and design, drafting of manuscript, critical revision. Gabriele Bonanno: Acquisition of data, critical revision. Omer Burak Demirel: Analysis and interpretation of data, critical revision. Steen Moeller: Analysis and interpretation of data, critical revision. Davide Piccini: Acquisition of data, critical revision. Burhaneddin Yaman: Analysis and interpretation of data, critical revision. Chetan Shenoy: Analysis and interpretation of data, critical revision. Matthias Stuber: Acquisition of data, critical revision. Chi Zhang: Study conception and design, drafting of manuscript, critical revision. Christopher W. Roy: Acquisition of data, critical revision.
Corresponding author
Ethics declarations
Conflict of interest
Davide Piccini and Gabriele Bonanno are employed by Siemens Healthineers AG, Bern, Switzerland. Matthias Stuber receives non-monetary research support from Siemens Healthineers that is covered by a master research agreement handled by his employer (CHUV). Matthias Stuber has a research contract with Circle that is handled by the Tech Transfer Office (PACTT) of his employer (CHUV).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, C., Piccini, D., Demirel, O.B. et al. Large-scale 3D non-Cartesian coronary MRI reconstruction using distributed memory-efficient physics-guided deep learning with limited training data. Magn Reson Mater Phy (2024). https://doi.org/10.1007/s10334-024-01157-8
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10334-024-01157-8