Abstract
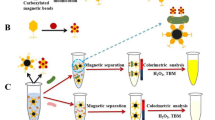
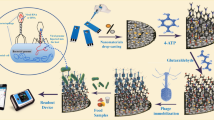
A broad host range phage-based nanozyme (Fe-MOF@SalmpYZU47) was prepared for colorimetric detection of multiple Salmonella enterica strains. The isolation of a broad host range phage (SalmpYZU47) capable of infecting multiple S. enterica strains was achieved. Then, it was directly immobilized onto the Fe-MOF to prepare Fe-MOF@SalmpYZU47, exhibiting peroxidase-like activity. The peroxidase-like activity can be specifically inhibited by multiple S. enterica strains, benefiting from the broad host range capture ability of Fe-MOF@SalmpYZU47. Based on it, a colorimetric detection approach was developed for S. enterica in the range from 1.0 × 102 to 1.0 × 108 CFU mL−1, achieving a low limit of detection (LOD) of 11 CFU mL−1. The Fe-MOF@SalmpYZU47 was utilized for detecting S. enterica in authentic food samples, achieving recoveries ranging from 91.88 to 105.34%. Hence, our proposed broad host range phage-based nanozyme exhibits significant potential for application in the colorimetric detection of pathogenic bacteria.
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Salmonella, a Gram-negative aerobic or partially anaerobic bacterium, is a prevalent zoonotic foodborne pathogen that can be transmitted through various food sources, including milk, raw vegetables, seafood, and livestock. Primarily, transmission occurs to humans through the consumption of animal-derived foods [1]. Human infection with Salmonella can result in a range of illnesses such as gastroenteritis, typhoid fever, septicemia, and potentially fatal outcomes in severe cases [2]. Salmonella is a genus of bacteria that includes over 2600 serotypes worldwide. One of the most common serotypes is Salmonella enterica (S. enterica), which comprises over 1500 serotypes and can cause human infection [3]. Outbreaks of foodborne illness are common due to the large number of S. enterica serotypes and their wide distribution. Therefore, the development of a rapid, accurate, sensitive, and reliable broad-spectrum detection method is imperative for precise identification and effective control of S. enterica contamination in food.
Currently, conventional methods for S. enterica detection encompass traditional plate culture colony counting [4], as well as molecular and immunological techniques such as polymerase chain reaction (PCR) [5] and enzyme-linked immunosorbent assay (ELISA) [6]. These techniques exhibit specificity and sensitivity; however, their drawbacks are apparent. The majority of them necessitate prolonged operation periods as well as specialized equipment and technicians, which can be expensive and impractical for on-site testing [7,8,9]. The increasing understanding of S. enterica has led to the development of a range of novel technologies in recent years, offering a wider array of options for detecting S. enterica. Optical sensors have gained significant momentum as an effective method for S. enterica detection, encompassing surface plasmon resonance (SPR) [10], fluorometry [11], colorimetry [12], and bioluminescence and chemiluminescence assays [13]. Among them, colorimetry exhibits immense potential due to its inherent simplicity, rapidity, cost-effectiveness, and ability to visually represent assay concentrations. Unfortunately, the sensitivity and specificity of colorimetric detection for S. enterica are commonly regarded as the primary obstacles [14].
Nanozymes, a class of nanomaterials possessing enzyme-like properties, exhibit several advantages over natural biological enzymes, including cost-effectiveness, facile synthesis, tunable catalytic activity, and exceptional stability [15,16,17]. Consequently, they can be employed as basic components in biosensors to augment the sensitivity of detection. For instance, Zhu et al. established a novel label-free ultrasensitive colorimetric aptamer nanozyme-based sensor for Staphylococcus aureus detection by immobilizing SA31 aptamer on the Mn3O4 nanozyme, with a LOD as low as 3 CFU mL−1 [18]. It can be seen that nanozyme can be combined with biorecognition units to specifically capture the target strains. However, these biorecognition units, such as aptamers and antibodies, suffer from poor stability, high probability of reaction inhibition, difficulty in preparation, and high costs to some extent [19, 20]. Therefore, it is necessary to develop a colorimetric method based on nanozymes combing with novel biorecognition elements for S. enterica detection.
Phages are viruses that specifically infect bacteria and always used as biorecognition elements in biosensors. They offer numerous advantages: (1) remarkable specificity towards host bacteria, (2) exceptional stability against harsh environmental conditions, and (3) outstanding affordability and fast multiplication in large quantities [21,22,23]. Given this, so much interesting work has been carried out based on phage-based biosensors [24, 25]. For example, Zhou et al. immobilized T2 phage on a glassy carbon electrode for electrochemical detection of Escherichia coli with high specificity [26]. The aforementioned study demonstrates the feasibility of employing phages as recognition elements in biosensors. However, a common limitation of these innovative assays is their ability to detect only a single serotype of bacteria, thus lacking broad-spectrum detection capabilities.
In this work, as shown in Scheme 1, a broad host range phage of S. enterica named SalmpYZU47 was isolated and purified. The biological characteristics of it were systematically investigated. After that, Fe-MOF was prepared using 2-aminoterephthalic acid (NH2-BDC) as a positively charged organic ligand. SalmpYZU47 was immobilized on the surface of Fe-MOF to form Fe-MOF@SalmpYZU47 by electrostatic interaction between SalmpYZU47 and NH2-BDC. The MOF@SalmpYZU47 obtained demonstrates peroxidase-like behavior by facilitating the chromogenic reaction of 3,3′,5,5′-tetramethylbenzidine (TMB) in the presence of hydrogen peroxide. Upon introduction of multiple S. enterica strains, the catalytic activity of it can be inhibited specifically, resulting in inhibition of TMB chromogenic reaction. Based on this principle, multiple S. enterica strains in food samples can be specifically, sensitively, and visually detected. In response to the current lack of research on the simultaneous detection of multiple strains in the food samples, Fe-MOF@SalmpYZU47 prepared in this study can detect multiple S. enterica strains due to the broad host spectrum property of SalmpYZU47, which greatly enhances the application potential of Fe-MOF@SalmpYZU47.
Experiment
Chemicals and reagents
Iron chloride hexahydrate (FeCl3·6H2O), 2-aminoterephthalic acid (NH2-BDC), sodium chloride (NaCl), and 3,3′,5,5′-tetramethylbenzidine (TMB) were sourced from Shanghai Aladdin Biochemical Technology Co., Ltd. Ethanol, hydrogen peroxide (H2O2), sodium acetate (NaAc), and acetic acid (HAc) were obtained from Sinopharm Chemical Reagent Co., Ltd. Pluronic F127 was purchased from Beyotime Biotechnology Co., Ltd. Luria–Bertani (LB) and agar powder medium were procured from Qingdao Hi-Tech Industrial Park Hope Bio-Technology Co., Ltd. All chemicals and reagents were used as received without further purification unless otherwise stated. Deionized water was utilized throughout the study.
Bacteria and phage culture
Thirty-two strains of S. enterica were used in this study (Table S1, (Supplementary data, SD)). Among them, 29 strains are sourced from the China Center of Industrial Culture Collection (CICC), and 3 strains were isolated from contaminated food samples. Staphylococcus aureus (S. aureus, CICC 21600), Escherichia coli (E. coli, CICC 10664), Bacillus cereus (B. cereus, CICC 21261), Pseudomonas fluorescens (P. fluorescens, ATCC 13525), Vibrio parahaemolyticus (V. parahaemolyticus, CICC 21617), and Listeria monocytogenes (L. monocytogenes, ATCC 19111) were obtained from the Laboratory of Prevention and Control of Biological Hazard Factors at Yangzhou University in Jiangsu Province, China. All bacteria were cultured at 37 °C in LB broth except for V. parahaemolyticus, which was cultured in LB broth supplemented with 3% (w/v) NaCl.
Phage was isolated from an effluent sample in Yangzhou using S. enterica CICC 21488 as the host bacterium. It was subsequently purified and propagated by employing the double-plate method, following previously described protocols [27]. The phage titer was quantified as plaque-forming units per milliliter (PFU mL−1). Finally, the resulting phage was named SalmpYZU47.
S. enterica phage experiments
Bacteriophage host range
The host range of SalmpYZU47 was determined by employing a spot test protocol, as previously described [27], using various S. enterica strains (Table S1, SD).
Optimal MOI determination
The concentration of the host bacteria was diluted to 106 CFU mL−1. Subsequently, a mixture of 0.1 mL phage and diluted bacterial solution was added to 5 mL Luria–Bertani (LB) broth in varying ratios: 100, 10, 1, 0.1, 0.01, 0.001, and 0.0001. The mixture was then incubated at 37 °C for 6 h. Following incubation, the culture underwent centrifugation at 8000 rpm for10 min and the resulting supernatant was filtered through a 0.22-µm filter. The optimal multiplicity of infection (MOI) was determined based on the highest titer observed in the filtrate.
Thermal stability and pH test
The phage suspension (0.5 mL) was incubated in a metal bath at temperatures ranging from 40 to 90 °C for different durations (20, 40, and 60 min) to determine the phage titers. Subsequently, the phage suspension (0.1 mL) was mixed with 0.9 mL SM buffer (0.05 M Tris, 0.1 M NaCl, 0.008 M MgSO4, pH 7.5) containing varying pH levels and incubated at 37 °C for 2 h before determining the titer.
One-step growth curve of phage
The one-step growth curves of the phage were generated at a MOI of 0.1 as previously described [28]. Phage growth parameters, including the latent period, burst time, and burst size, were estimated based on these growth curves.
Adsorption rate of phage
The determination of phage adsorption was conducted according to previously reported method with minor modifications [27]. Briefly, 1 mL SM medium containing host cells and phages was incubated for 15 min, followed by centrifugation at 12,000 rpm for 30 min to remove bacteria and bound phages. The quantification of unabsorbed phages in the supernatants was then performed using the double-plate method.
Genome sequencing, assembly, and annotation
Genomic DNA of SalmpYZU47 was extracted using the ABigen Lambda Phage DNA Purification Kit (AB1141, ABigen Corporation, Beijing, China). Subsequently, the DNA extracts that met the sequencing requirements were submitted to Shanghai Lingen Biotechnology Co., Ltd. (Shanghai, China) for whole genome sequencing. Gene models were identified using GeneMark. All gene models were then subjected to functional annotation by comparing them against the non-redundant (NR in NCBI) database, Swiss-Prot (http://uniprot.org), KEGG (http://www.genome.jp/kegg/), and COG (http://www.ncbi.nlm.nih.gov/COG) through blastp module. Finally, a linear representation of the annotated predicted ORFs was generated using IBS 1.0.
Preparation of Fe-MOF and Fe-MOF@SalmpYZU47
First, 0.32 g Pluronic F127 was dissolved in 26.68 mL deionized water. Next, 3.32 mL FeCl3·6H2O aqueous solution (0.66 mmol) was added and stirred for 1 h. 0.3 mL acetic acid was added and stirred for 1 h, followed by the addition of 120 mg NH2-BDC and stirring for another 2 h. The resulting mixture was transferred to tetrafluoroethylene-lined autoclave at 110 °C for 24 h. The dark brown solid Fe-MOF was collected by centrifugation and washed three times with alternating water and ethanol. After that, it was dried under vacuum at 60 °C. Finally, 15 mg Fe-MOF was dissolved in 4 mL deionized water and then mixed with 4 mL phage SalmpYZU47 (8.0 × 109 PFU mL−1). The residual of SalmpYZU47 was first removed by centrifugation and then washed by deionized water. The final precipitate is then resuspended in 8 mL of deionized water, thereby yielding the prepared Fe-MOF@SalmpYZU47. The obtained Fe-MOF@SalmpYZU47 was stored at 4 °C for further use.
Characterization
The microstructures were observed using a transmission electron microscopy (TEM, Tecnai 12, Netherlands). Crystal structure measurements were conducted utilizing an X-ray diffractometer (XRD, D8 Advance, Bruker AXS, Germany). Quantitative elemental analyses were performed employing an X-ray electron spectrometer (XPS, ESCALAB 250Xi, Thermo Scientific, America). Free radicals were detected via electron paramagnetic resonance (EPR, A300-10/12, Bruker, Germany).
Peroxidase-like activity of Fe-MOF@SalmpYZU47
Peroxidase-like activity of Fe-MOF@SalmpYZU47 was evaluated through TMB chromogenic reaction. Briefly, 100 µL Fe-MOF@SalmpYZU47, 100 µL H2O2 (100 mM), and 100 µL TMB (5 mM, dissolved in ethanol) were sequentially added into 2700 µL NaAc-HAc buffer solution (0.2 M, pH 4.0). The mixture was then incubated for 30 min. The UV–vis spectrophotometer (UV-3200, Mapada, China) was used to measure the absorbance immediately.
Colorimetric detection of S. enterica using Fe-MOF@SalmpYZU47
One hundred microliters of Fe-MOF@SalmpYZU47 and 100 µL of S. enterica with varying concentrations (1.0 × 102 ~ 1.0 × 108 CFU mL−1) were added into 2600 µL NaAc-HAc buffer solution (0.2 M, pH 4.0), incubating for 10 min. Then, 100 µL H2O2 (100 mM) and 100 µL TMB (5 mM, dissolved in ethanol) were added, incubating for another 30 min. The absorbance was measured using a UV–vis spectrophotometer immediately.
Specificity and anti-interference tests
The specificity and anti-interference properties of Fe-MOF@SalmpYZU47 + TMB reaction system were investigated. The possible co-existing strains included S. aureus, E. coli, B. cereus, P. fluorescens, V. parahaemolyticus, and L. monocytogenes. Dead S. enterica was obtained from living S. enterica incubating at 100 °C for 15 min.
Specificity test
One hundred microliters of Fe-MOF@SalmpYZU47 and 100 µL of bacteria were added into 2600 µL NaAc-HAc buffer solution (0.2 M, pH 4.0), incubating for 10 min. Then, 100 µL H2O2 (100 mM) and 100 µL TMB (5 mM, dissolved in ethanol) were added, incubating for another 30 min. The absorbance was measured using a UV–vis spectrophotometer immediately.
Anti-interference test
One hundred microliters of other bacteria, 100 µL of S. enterica, and 100 µL of Fe-MOF@SalmpYZU47 were added into 2500 µL NaAc-Hac buffer solution (0.2 M, pH 4.0), incubating for 10 min. Then, 100 µL H2O2 (100 mM) and 100 µL TMB (5 mM, dissolved in ethanol) were added, incubating for another 30 min. The absorbance was measured using a UV–vis spectrophotometer immediately.
Authentic food sample pretreatment and detection
Milk, bass, and chicken were selected as real food samples for verifying the feasibility of Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system. The samples were purchased from local supermarkets in Yangzhou. Bass and chicken were processed into broth by boiling them with drinking water at a mass ratio of 1:10. The meat residue was filtered through gauze to obtain the broth, which was then sterilized by autoclaving at 121 °C for 20 min to obtain sterile broth. S. enterica with known concentration (3 × 102, 3 × 104, and 3 × 106 CFU mL−1) was added into the prepared samples. Subsequently, both the Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system and traditional colony counting method were employed to determine these samples.
Results and discussion
Characterization of SalmpYZU47
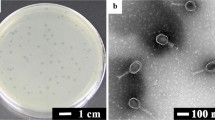
SalmpYZU47 were isolated from sewage samples, and its biological characteristics were investigated. As shown in Fig. 1A, SalmpYZU47 was observed to be classified within the family Myoviridae. According to Table S1 (SD), it remained excellent lysis of 15 other S. enterica strains. As depicted in Fig. 1B, the optimal MOI of SalmpYZU47 was determined to be 0.001. At this optimum MOI, the viral titer reached up to 8.5 × 109 PFU mL−1. Moreover, the titer of SalmpYZU47 was consistently maintained at a high level of 1.6 × 109 PFU mL−1 across a wide pH ranging from 3.0 to 11.0 (Fig. 1C), while it exhibited remarkable stability below 70 °C (Fig. 1D). In addition, SalmpYZU47 exhibited a maximum adsorption rate of 54.1% within a 10 min adsorption period (Fig. 1E). Furthermore, its one-step growth curve displayed a latency period of 10 min and a burst size of 220 PFU cell−1 (Fig. 1F). These physiological properties exhibited by SalmpYZU47 indicate its significant potential as a biosensor recognition unit due to its resistance to acid, alkali, and heat, rapid adsorption capability, and large burst size.
TEM image (A), phage titer (B), pH (C), thermal sensitivity (D), adsorption rate (E), one-step growth curve (L, latent period; R, rise period; P, plateau period) (F), and mosaic structure (positions, orientations, and arrows with predicted functions represent the functions of predicted ORFs) (G) of SalmpYZU47
Genome sequencing results of SalmpYZU47 revealed a genome size of 85,856 bp, displaying a GC content of 50.23%. The analysis identified a total of 126 open reading frames (ORFs), with 45 encoding known proteins and 81 encoding hypothetical proteins (Fig. 1G). The blast comparison analysis confirmed that SalmpYZU47 belongs to the Myoviridae family, Felixounavirus genus. SalmpYZU47 possesses a contractile tail sheath along with both long and short tail fibers, accompanied by a baseplate. The tail exhibits high specificity towards receptors on the bacterial cell, thereby facilitating efficient delivery of genetic material within the host cell [8]. The adsorption and infection of SalmpYZU47 to the host are determined by the tail structure and its RBP [29], thereby indicating specific recognition and adsorption features of SalmpYZU47 through tail fibronectin towards S. enterica, rendering it an ideal foundation for targeted detection.
Synthesis and characterization of Fe-MOF@SalmpYZU47
The synthesis of Fe-MOF was successfully accomplished using a solvothermal method [30]. First, as depicted in Fig. 2B, XRD was used to analyze the crystal structure of the obtained sample. It exhibited characteristic diffraction peaks, which are in complete agreement with the distinctive diffraction peaks of NH2-MIL-88B(Fe) according to the previously reported method [30]. Hence, our proposed Fe-MOF essentially is NH2-MIL-88B(Fe). Then, as shown in Fig. 2C, Fe-MOF exhibits a spindle-shaped nanoparticle measuring approximately 300 nm in length and 100 nm in width. The XPS was adopted to explore the surface chemical state and structural details of Fe-MOF. As displayed in Fig. 2E, Fe-MOF consists of element carbon (C 1s), nitrogen (N 1s), oxygen (O 1s), and iron (Fe 2p). Furthermore, the analytical spectrum of Fe 2p valence state exhibits two prominent peaks and a distinct satellite. Specifically, the Fe 2p 3/2 and Fe 2p 1/2 binding energies were determined to be 710.7 eV and 724.4 eV, respectively (Fig. 2F). These results indicate successful synthesis of Fe-MOF.
SalmpYZU47 was immobilized on the surface of Fe-MOF through electrostatic interaction (Fig. 2A). As depicted in Fig. 2D, the head of SalmpYZU47 was securely affixed to the Fe-MOF surface, while the tail protruded outward, thereby preserving its inherent biological structure. The XRD pattern revealed that Fe-MOF@SalmpYZU47 exhibited identical peak positions as Fe-MOF, albeit with a gradual decrease in intensity, implying the preservation of the Fe-MOF structure upon adherence to SalmpYZU47 (Fig. 2B). Furthermore, XPS revealed that Fe-MOF@SalmpYZU47 contained element sulfur (S 2p), indicating successful binding of SalmpYZU47 on the surface of Fe-MOF (Fig. 2E). The biological activity of SalmpYZU47 after immobilization was verified through spotting experiments, which demonstrated that SalmpYZU47 maintained its remarkable lysogenic activity (Fig. 2G).
Peroxidase-like activity of Fe-MOF@SalmpYZU47
Peroxidase-like activity of Fe-MOF@SalmpYZU47 was studied. The absorption peak at 625 nm of Fe-MOF + H2O2 + TMB reaction system is greater than that of the Fe-MOF@SalmpYZU47 + H2O2 + TMB system (Fig. S1, SD). This indicates that the immobilization of SalmpYZU47 decreases peroxidase-like activity of Fe-MOF. As shown in Fig. 3A, Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system had a distinct absorption peak at 652 nm, which was attributed to the generation of blue oxidized TMB (TMBox), while the other three systems did not show significant color change. This result indicates that Fe-MOF@SalmpYZU47 has the ability to expedite the TMB chromogenic reaction in the presence of H2O2. To further investigate the mechanism, the free radicals generated during the TMB chromogenic reaction were first captured using DMPO and then detected using EPR. As depicted in Fig. 3B, the distinctive peaks observed in the generated free radicals corresponded to those of hydroxyl radicals (·OH). Thus, this demonstrated that Fe-MOF@SalmpYZU47 exhibits peroxidase-like activity, facilitating the breakdown of H2O2 to produce hydroxyl radicals, which can oxidize TMB to form blue TMBox.
UV–vis spectra of four reaction systems (inset is the corresponding image) (A) and EPR spectrum of Fe-MOF@SalmpYZU47 + H2O2 + DMPO reaction system (inset is the corresponding reaction mechanism of Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system) (B), effect of buffer solution pH (C) and Fe-MOF@SalmpYZU47 amount (D) on the Fe-MOF@SalmpYZU47 + H2O2 + TMB chromogenic system, and steady-state catalytic kinetics of Fe-MOF@SalmpYZU47 using TMB (E) and H2O2 (F) as substrates
The Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system was optimized to achieve the best property. As shown in Fig. 3C, the catalytic activity of Fe-MOF@SalmpYZU47 exhibited an initial increase followed by a subsequent decrease in response to variations in the pH of NaAc-Hac buffer. Notably, the highest level of catalytic activity of Fe-MOF@SalmpYZU47 was observed at pH 4.0, which consequently emerged as the optimal pH for subsequent experimentation. Then, the effect of Fe-MOF@SalmpYZU47 amount on the reaction system was investigated. As shown in Fig. 3D, the absorbance at 652 nm of reaction system exhibited an increase with the amount increment. However, taking into account both cost and overall effectiveness, 100 µL Fe-MOF@SalmpYZU47 was selected as the optimal parameter in the following experiments. The absorbance at 652 nm of the Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system over time was monitored. As depicted in Fig. S2 (SD), a positive correlation between time and the absorbance at 652 nm of the reaction system was observed. Moreover, increasing the incubation time for a short period did not impede reaction system, and a significant color reaction was already evident after 30 min. Therefore, to optimize incubation time, 30 min was set as the best parameter for subsequent experiments. Moreover, the robustness of the Fe-MOF against harsh incubation pH, temperature, salt, and time was also investigated. As shown in Fig. S3 (SD), the robustness of the Fe-MOF against harsh environment is perfect. This lays the foundation of wide application potential of Fe-MOF@SalmpYZU47.
Additionally, steady-state kinetic measurements were carried out using H2O2 and TMB as substrates. As displayed in Fig. 3E, F, the measured data was fitted by Michaelis–Menten equation. The Km were calculated as 1.0 and 1.2 mM using H2O2 and TMB as substrates, respectively. And the Vmax were calculated as 5.8 and 25.3 × 10−8 M s−1 using H2O2 and TMB as substrates, respectively. Compared with previously reported peroxidase-like nanozymes (Table S2, SD) [31,32,33,34], it is seen that Fe-MOF@SalmpYZU47 possesses remarkable affinity and catalytic efficiency towards TMB and H2O2. Hence, Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system can serve as a robust foundation for the current targeted assay.
Colorimetric detection of S. enterica using Fe-MOF@SalmpYZU47
The strict host specificity observed between SalmpYZU47 and S. enterica renders Fe-MOF@SalmpYZU47 highly promising for the specific detection of S. enterica. As shown in Fig. 4A, the absorbance at 652 nm of Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system exhibited a significant decrease upon the addition of S. enterica (CICC 21488), which was visually discernible to the naked eye. This phenomenon suggests that S. enterica (CICC 21488) can inhibit the Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system. In addition, 15 other S. enterica strains were also employed due to their successful spotting experiments. As displayed in Fig. 4B, it is found that 15 other S. enterica strains can also obviously inhibit Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system. And the inhibitory effect of 15 other S. enterica strains is almost same as that of S. enterica (CICC 21488), indicating that Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system can be applied for detection of multiple S. enterica strains. Moreover, the inhibitory effect of living/dead S. enterica towards Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system was also studied. As depicted in Fig. S4 (SD), both living and dead S. enterica can inhibit the reaction system. Specific biological interaction exists between the phage and its host bacteria. The specificity and selectivity of phage attachment and infection of host cells is determined not only by the phage recognition site but also by the structure and receptors on the bacterial cell surface. Although host cells are no longer alive, the receptor proteins on its surface are consistent with those of living host cells. The unique tail protein gene in the phage genome can specifically and recognize and capture living/dead S. enterica [33, 34]. This feature of Fe-MOF@SalmpYZU47 + H2O2 + TMB chromogenic system can alert food safety after S. enterica contamination, whether target bacteria are alive or dead. Based on the experimental results, it can be inferred that Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system can be employed to detect a series of S. enterica strains, which profits from a broad host spectrum of SalmpYZU47.
UV–vis spectra of Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system with/without S. enterica (CICC 21488) (inset is the corresponding real image) (A), the inhibitory effect of 15 other S. enterica strains towards Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system (inset is the corresponding real image) (B), TEM images of S. enterica (C), Fe-MOF@SalmpYZU47 and S. enterica composites (D, E), and colorimetric detection mechanism of S. enterica using Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system (F)
To further investigate the internal detection mechanism of Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system towards S. enterica, the micromorphology of S. enterica captured by MOF@SalmpYZU47 was obtained. As shown in Fig. 4C, S. enterica has an intact cell structure. Upon addition of Fe-MOF@SalmpYZU47, S. enterica was tightly surrounded by MOF@SalmpYZU47 (Fig. 4D). And the tails of SalmpYZU47 on the surface of Fe-MOF@SalmpYZU47 capture S. enterica tightly (Fig. 4E). As shown in Fig. 4F, it can be concluded that MOF@SalmpYZU47 can specifically capture S. enterica, first resulting in the blocks of enzyme-like catalytic sites of Fe-MOF@SalmpYZU47 and then indirectly inhibiting TMB chromogenic reaction. This specific capture ability of Fe-MOF@SalmpYZU47 originates from the inherent genes of SalmpYZU47. It is well known that caudovirus phages possess the capacity to infect a diverse range of hosts attributed to their distinctive tail structure and recognition sites [35]. According to genomic analysis, the tail fibronectin in SalmpYZU47 possesses abundant distinctive recognition sites, conferring Fe-MOF@SalmpYZU47 with a specific recognition capability.
For high-performance detection of S. enterica, Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system was further optimized. The incubation time between Fe-MOF@SalmpYZU47 and S. enterica is shown in Fig. S5 (SD). The absorbance at 652 nm exhibited a gradual decrease over time and reached a steady state after 10 min of incubation. Consequently, an incubation period of 10 min was determined as the optimal duration for the interaction between Fe-MOF@SalmpYZU47 and S. enterica. After that, the absorbance change at 652 nm of Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system upon addition of various concentrations of S. enterica was studied. As depicted in Fig. 5A, the absorbance at 652 nm of Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system exhibited a decrease with increasing concentration of S. enterica. As displayed in Fig. 5B, the absorbance at 652 nm of Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system possesses a linear relationship with the logarithm of S. enterica level (1.0 × 102 ~ 1.0 × 108 CFU mL−1), providing an equation: Y = − 0.13X + 1.30 (R2 = 0.987). Furthermore, the detection limit (LOD) is determined to be as low as 11 CFU mL−1 according to S/N = 3. Moreover, the specificity and anti-interference of Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system were verified. S. aureus, E. coli, B. cereus, P. fluorescens, V. parahaemolyticus, and L. monocytogenes were set as the possible co-existing disturbing bacteria. As shown in Fig. 5C, other bacteria almost have no significant effect on the Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system, highlighting its high specificity. Then, Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system exhibits remarkable accuracy even in the presence of other bacteria, suggesting its strong high anti-interference capability (Fig. 5D). These characteristics ensure the potential application of the Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system in the detection of complex samples.
UV–vis spectra of Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system with different concentrations of S. enterica (A), linear fitting between absorbance at 652 nm of reaction system and the logarithm of S. enterica concentration (B), and specificity (C) and anti-interference (D) of Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system
As listed in Table S3 (SD), compared to other detection methods, such as ELISA [36], electrochemical immunosensor [37], fiber optic surface plasmon resonance [38], and fluorimetry [39], our proposed Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system offers several advantages for S. enterica detection: (1) visualization, (2) wide detection range, (3) specificity, and (4) convenience. These advantages are primarily attributed to high peroxidase-like activity of Fe-MOF and excellent specificity of SalmpYZU47 towards S. enterica. Moreover, the high sensitivity of Fe-MOF@SalmpYZU47 + H2O2 + TMB chromogenic system towards S. enterica is attributed to the signal amplification of TMB chromogenic system catalyzed by Fe-MOF@SalmpYZU47 nanozyme.
To evaluate the practical detection ability of the Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system, S. enterica in milk, bass, and chicken was detected. The results, shown in Table S4 (SD), indicated good detection performance with recoveries ranging from 91.88 to 105.34%. This result indicates that our proposed Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system can be employed for practical applications of S. enterica detection in food samples.
Conclusion
In summary, a broad host range phage SalmpYZU47, which can specifically lyse multiple S. enterica strains, was isolated and purified. Then, the phage SalmpYZU47 was immobilized on the surface of Fe-MOF via electrostatic interaction. The Fe-MOF@SalmpYZU47 obtained exhibits remarkable peroxidase-like activity, being able to catalyze TMB chromogenic reaction in presence of H2O2. The Fe-MOF@SalmpYZU47 + H2O2 + TMB reaction system can be employed to detect multiple S. enterica strains with high sensitivity, specify, and anti-interfere. This is main attributed to high peroxidase-like activity of Fe-MOF and a broad host range of SalmpYZU47 towards multiple S. enterica strains. In addition, the detection in real food samples achieves an excellent effect. These results show great potential of Fe-MOF@SalmpYZU47 as a general tool for detecting pathogenic bacteria in practice.
References
Bearson S (2022) Salmonella in swine: prevalence, multidrug resistance, and vaccination strategies. Annu Rev Animal Biosci 10:373–393. https://doi.org/10.1146/annurev-animal-013120-043304
Guan L, Hu A, Ma S, Liu J, Yao X, Ye T, Han M, Yang C, Zhang R, Xiao X, Wu Y (2024) Lactiplantibacillus plantarum postbiotic protects against Salmonella infection in broilers via modulating NLRP3 inflammasome and gut microbiota. Poult Sci 103(4):103483–103483. https://doi.org/10.1016/j.psj.2024.103483
Jennings E, Thurston T, Holden D (2017) Salmonella SPI-2 type III secretion system effectors: molecular mechanisms and physiological consequences. Cell Host Microbe 22(2):217–231. https://doi.org/10.1016/j.chom.2017.07.009
Collier S, Deng L, Adam E, Benedict K, Beshearse E, Blackstock A, Bruce B, Derado G, Edens C, Fullerton K, Gargano J, Geissler A, Hall A, Havelaar A, Hill V, Hoekstra R, Reddy S, Scallan E, Stokes E, Yoder J, Beach M (2021) Estimate of burden and direct healthcare cost of infectious waterborne disease in the United States. Emerg Infect Dis 27(1):140–149. https://doi.org/10.3201/eid2701.190676
Vinayaka A, Ngo T, Kant K, Engelsmann P, Dave V, Shahbazi M, Wolff A, Bang D (2019) Rapid detection of Salmonella enterica in food samples by a novel approach with combination of sample concentration and direct PCR. Biosens Bioelectron 129:224–230. https://doi.org/10.1016/j.bios.2018.09.078
Jiao C, Duan W, Wu X, Shang Y, Zhang F, Zhang M, Chen X, Zeng J, Yang C (2023) Multifunctional nanoprobe-amplified enzyme-linked immunosorbent assay on capillary: a universal platform for simple, rapid, and ultrasensitive dual-mode pathogen detection. Anal Chem 95(30):11316–11325. https://doi.org/10.1021/acs.analchem.3c01375
Castle L, Schuh D, Reynolds E, Furst A (2021) Electrochemical sensors to detect bacterial foodborne pathogens. Acs Sensors 6(5):1717–1730. https://doi.org/10.1021/acssensors.1c00481
Hussain W, Ullah M, Farooq U, Aziz A, Wang S (2021) Bacteriophage-based advanced bacterial detection: concept, mechanisms, and applications. Biosens Bioelectron 177:112973–112973. https://doi.org/10.1016/j.bios.2021.112973
Qin L, Xiao J, Yang H, Liang J, Li L, Wu S, Peng D (2024) Rapid immunoassays for the detection of quinoxalines and their metabolites residues in animal-derived foods: a review. Food Chem 443:138539–138539. https://doi.org/10.1016/j.foodchem.2024.138539
Ma S, Luo X, Kong J, Li X, Cao Z, Wang X, Cai W, Wang L, Ran G (2022) Plasmonic silver loaded hybrid Bi-Ag nanoalloys for highly efficient disinfection by enhancing photothermal performance and interface capability. Chem Eng J 450:138016. https://doi.org/10.1016/j.cej.2022.138016
Zhao J, Han M, Ma A, Jiang F, Chen R, Dong Y, Wang X, Ruan S, Chen Y (2024) A machine vision-assisted Argonaute-mediated fluorescence biosensor for the detection of viable Salmonella in food without convoluted DNA extraction and amplification procedures. J Hazard Mater 466:133648–133648. https://doi.org/10.1016/j.jhazmat.2024.133648
Qi W, Zheng L, Hou Y, Duan H, Wang L, Wang S, Liu Y, Li Y, Liao M, Lin J (2022) A finger-actuated microfluidic biosensor for colorimetric detection of foodborne pathogens. Food Chem 381:131801–131801. https://doi.org/10.1016/j.foodchem.2021.131801
Lin X, Zhao M, Peng T, Zhang P, Shen R, Jia Y (2023) Detection and discrimination of pathogenic bacteria with nanomaterials-based optical biosensors: a review. Food Chem 426:136578–136578. https://doi.org/10.1016/j.foodchem.2023.136578
Zhang X, Lin S, Liu S, Tan X, Dai Y, Xia F (2021) Advances in organometallic/organic nanozymes and their applications. Coord Chem Rev 429:213652. https://doi.org/10.1016/j.ccr.2020.213652
Feng M, Li X, Zhang X, Huang Y (2023) Recent advances in the development and analytical applications of oxidase-like nanozymes. Trac-Trends Anal Chem 166:117220. https://doi.org/10.1016/j.trac.2023.117220
Niu K, Chen J, Lu X (2023) Versatile biomimetic catalyst functionalized nanozymes for electrochemical sensing. Chem Eng J 475:146491. https://doi.org/10.1016/j.cej.2023.146491
Zhao Y, Wang X, Pan S, Hong F, Lu P, Hu X, Jiang F, Wu L, Chen Y (2024) Bimetallic nanozyme-bioenzyme hybrid material-mediated ultrasensitive and automatic immunoassay for the detection of aflatoxin B1 in food. Biosens Bioelectron 248:115992. https://doi.org/10.1016/j.bios.2023.115992
Zhu S, Tang Y, Shi B, Zou W, Wang X, Wang C, Wu Y (2021) Oligonucleotide-mediated the oxidase-mimicking activity of Mn3O4 nanoparticles as a novel colorimetric aptasensor for ultrasensitive and selective detection of Staphylococcus aureus in food. Sens Actuators B-Chem 349:130809. https://doi.org/10.1016/j.snb.2021.130809
Costa S, Nogueira C, Cunha A, Lisac A, Carvalho C (2023) Potential of bacteriophage proteins as recognition molecules for pathogen detection. Crit Rev Biotechnol 43(5):787–804. https://doi.org/10.1080/07388551.2022.2071671
Xu X, Xu Q, Li W, Xiao F, Xu H (2024) From engineered photoactive materials to detection signal amplification strategies in photoelectrochemical biosensing of pathogens: new horizons and perspectives. Chem Eng J 480:147941. https://doi.org/10.1016/j.cej.2023.147941
Hatfull G, Dedrick R, Schooley R (2022) Phage therapy for antibiotic-resistant bacterial infections. Annu Rev Med 73:197–211. https://doi.org/10.1146/annurev-med-080219-122208
Strathdee S, Hatfull G, Mutalik V, Schooley R (2023) Phage therapy: from biological mechanisms to future directions. Cell 186(1):17–31. https://doi.org/10.1016/j.cell.2022.11.017
Uyttebroek S, Chen B, Onsea J, Ruythooren F, Debaveye Y, Devolder D, Spriet I, Depypere M, Wagemans J, Lavigne R, Pirnay J, Merabishvili M, Munter P, Peetermans W, Dupont L, Gerven L, Metsemakers W (2022) Safety and efficacy of phage therapy in difficult-to-treat infections: a systematic review. Lancet Infect Dis 22(8):E208–E220. https://doi.org/10.1016/s1473-3099(21)00612-5
Huang C, Zhao J, Lu R, Wang J, Nugen S, Chen Y, Wang X (2023) A phage-based magnetic relaxation switching biosensor using bioorthogonal reaction signal amplification for Salmonella detection in foods. Food Chem 400:134035–134035. https://doi.org/10.1016/j.foodchem.2022.134035
Ye J, Guo J, Li T, Tian J, Yu M, Wang X, Majeed U, Song W, Xiao J, Luo Y, Yue T (2022) Phage-based technologies for highly sensitive luminescent detection of foodborne pathogens and microbial toxins: a review. Compr Rev Food Sci Food Safety 21(2):1843–1867. https://doi.org/10.1111/1541-4337.12908
Zhou Y, Marar A, Kner P, Ramasamy R (2017) Charge-directed immobilization of bacteriophage on nanostructured electrode for whole-cell electrochemical biosensors. Anal Chem 89(11):5735–5742. https://doi.org/10.1021/acs.analchem.6b03751
Gao L, Ouyang M, Li Y, Zhang H, Zheng X, Li H, Rao S, Yang Z, Gao S (2022) Isolation and characterization of a lytic vibriophage OY1 and its biocontrol effects against Vibrio spp. Front Microbiol 13:830692–830692. https://doi.org/10.3389/fmicb.2022.830692
Rai S, Tyagi A, Kumar B, Reddy S (2023) Isolation and characterization of Aeromonas hydrophila lytic phage, and evaluation of a phage cocktail against A. hydrophila contamination in fish fillet. Food Control 145:109460. https://doi.org/10.1016/j.foodcont.2022.109460
Denyes J, Dunne M, Steiner S, Mittelviefhaus M, Weiss A, Schmidt H, Klumpp J, Loessner M (2017) Modified bacteriophage S16 long tail fiber proteins for rapid and specific immobilization and detection of Salmonella cells. Appl Environ Microbiol 83(12):E00277-E317. https://doi.org/10.1128/aem.00277-17
Wang X, Yun Y, Sun W, Lu Z, Tao X (2022) A high-performance fluorescence immunoassay based on pyrophosphate-induced MOFs NH2-MIL-88B(Fe) hydrolysis for chloramphenicol detection. Sens Actuators B-Chem 353:131143. https://doi.org/10.1016/j.snb.2021.131143
Darabdhara G, Sharma B, Das M, Boukherroub R, Szunerits S (2017) Cu-Ag bimetallic nanoparticles on reduced graphene oxide nanosheets as peroxidase mimic for glucose and ascorbic acid detection. Sens Actuators B-Chem 238:842–851. https://doi.org/10.1016/j.snb.2016.07.106
Wei J, Chen X, Shi S, Mo S, Zheng N (2015) An investigation of the mimetic enzyme activity of two-dimensional Pd-based nanostructures. Nanoscale 7(45):19018–19026. https://doi.org/10.1039/c5nr05675f
Zhang Y, Xu X, Yang J, Tan M, Zhou W, Ga L, Yang Z (2023) Directional immobilization of phage on the palladium-based nanozyme for colorimetric detection of Cronobacter sakazakii in powdered infant formula. LWT-Food Sci Technol 186:115260. https://doi.org/10.1016/j.lwt.2023.115260
Zhou W, Wen H, Hao G, Zhang Y, Yang J, Gao L, Zhu G, Yang Z, Xu X (2023) Surface engineering of magnetic peroxidase mimic using bacteriophage for high-sensitivity/specificity colorimetric determination of Staphylococcus aureus in food. Food Chem 426:136611–136611. https://doi.org/10.1016/j.foodchem.2023.136611
Arnaud C, Effantin G, Vives C, Engilberge S, Bacia M, Boulanger P, Girard E, Schoehn G, Breyton C (2017) Bacteriophage T5 tail tube structure suggests a trigger mechanism for Siphoviridae DNA ejection. Nat Commun 8:1953. https://doi.org/10.1038/s41467-017-02049-3
Ren Y, Wei J, Wang Y, Wang P, Ji Y, Liu B, Wang J, González-Sapienza G, Wang Y (2022) Development of a streptavidin-bridged enhanced sandwich ELISA based on self-paired nanobodies for monitoring multiplex Salmonella serogroups. Anal Chim Acta 1203:339705–339705. https://doi.org/10.1016/j.aca.2022.339705
Feng K, Li T, Ye C, Gao X, Yang T, Liang X, Yue X, Ding S, Dong Q, Yang M, Xiong C, Huang G, Zhang J (2021) A label-free electrochemical immunosensor for rapid detection of Salmonella in milk by using CoFe-MOFs-graphene modified electrode. Food Control 130:108357. https://doi.org/10.1016/j.foodcont.2021.108357
Zhou C, Zou H, Li M, Sun C, Ren D, Li Y (2018) Fiber optic surface plasmon resonance sensor for detection of E. coli O157:H7 based on antimicrobial peptides and AgNPs-rGO. Biosens Bioelectron 117:347–353. https://doi.org/10.1016/j.bios.2018.06.005
Yang M, Chen X, Zhu L, Lin S, Li C, Li X, Huang K, Xu W (2021) Aptamer-functionalized DNA-silver nanocluster nanofilm for visual detection and elimination of bacteria. ACS Appl Mater Interfaces 13(32):38647–38655. https://doi.org/10.1021/acsami.1c05751
Funding
This study is supported by the Natural Science Foundation of the Open Research Fund of Key Laboratory of Healthy Freshwater Aquaculture, Ministry of Agriculture and Rural Affairs (ZJK202314), the Natural Science Foundation of Jiangsu Province (BK20230586), the Natural Science Foundation of Yangzhou City (SZR2023000023), and the Higher Education Institutions of Jiangsu Province (23KJB550011).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Ethics approval
This research did not involve human or animal samples.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gao, L., Zhang, L., Yang, J. et al. Immobilization of a broad host range phage on the peroxidase-like Fe-MOF for colorimetric determination of multiple Salmonella enterica strains in food. Microchim Acta 191, 331 (2024). https://doi.org/10.1007/s00604-024-06402-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-024-06402-4